The focus of the Pinello laboratory is to use innovative computational approaches and cutting-edge experimental assays such as CRISPR genome editing and single cell sequencing to systematically analyze sources of genetic and epigenetic variation and gene expression variability in human traits and diseases.
The lab uses machine learning, data mining and high performance computing technologies, for instance parallel computing and the new cloud oriented architectures, to solve computationally challenging and Big Data problems associated with next generation sequencing data analysis.
Our mission is to use computational strategies to further our understanding of disease etiology and to provide a foundation for the development of new drugs and more targeted treatments.
Research Interests
(epi)Genomics
We develop computational approaches to investigate different epigenetic factors that regulate the chromatin structure and the gene expression integrating NGS for instance: ChIP-seq, RNA-seq, ATAC-seq and Bisulfite-seq.

Haystack Analysis on H3K27ac for 6 Cell Lines
Single Cell Analysis
We have experience analyzing data from recent single cell assays for gene expression and chromatin accessibility, such as scRNA-seq and scATAC-seq.
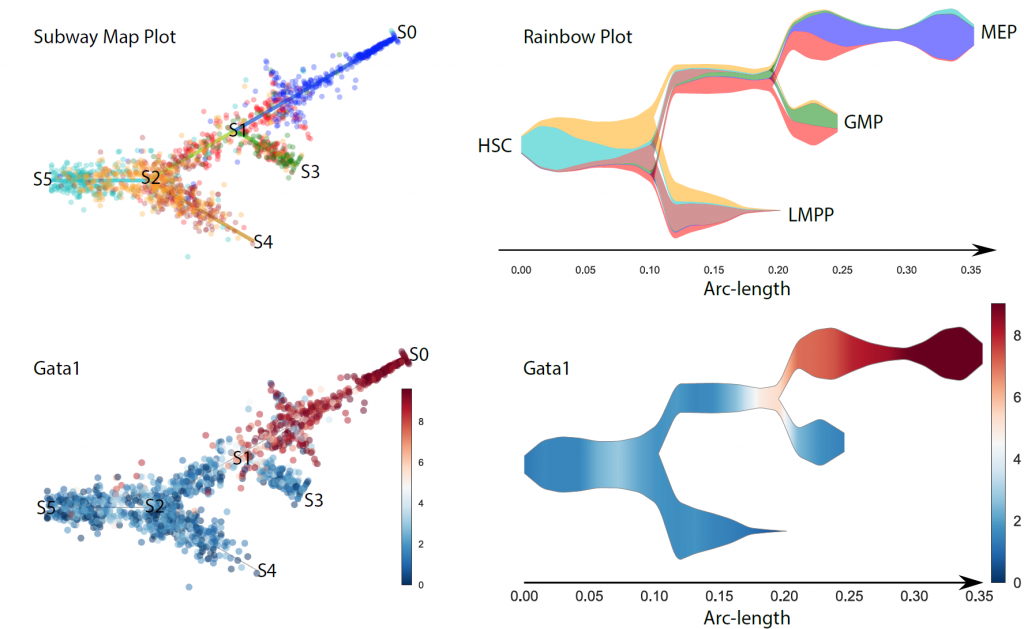
STREAM Trajectory Analysis on Mouse Haematopoiesis
Genome Editing
We embraced the new revolution in functional genomics made possible by the novel genome editing approaches such as CRISPR/Cas9, base editor and prime editors developing new tools to design gene editing experiments and quantify and visualize edit outcomes from sequencing data.
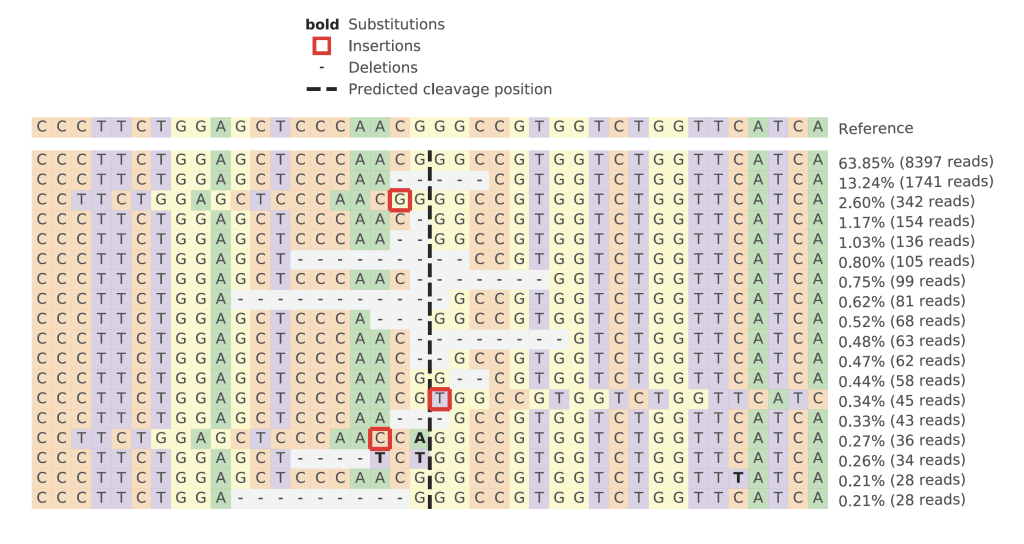
Crispresso2 Analysis on Targeted Amplicon Sequencing
